RESEARCH ARTICLE
Retrieval of Radiology Reports Citing Critical Findings with Disease-Specific Customization
Ronilda Lacson*, Nathanael Sugarbaker, Luciano M Prevedello, IP Ivan, Wendy Mar, Katherine P Andriole, Ramin Khorasani
Article Information
Identifiers and Pagination:
Year: 2012Volume: 6
First Page: 28
Last Page: 35
Publisher Id: TOMINFOJ-6-28
DOI: 10.2174/1874431101206010028
Article History:
Received Date: 15/5/2012Revision Received Date: 29/6/2012
Acceptance Date: 14/7/2012
Electronic publication date: 10/8/2012
Collection year: 2012
open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
Abstract
Background:
Communication of critical results from diagnostic procedures between caregivers is a Joint Commission national patient safety goal. Evaluating critical result communication often requires manual analysis of voluminous data, especially when reviewing unstructured textual results of radiologic findings. Information retrieval (IR) tools can facilitate this process by enabling automated retrieval of radiology reports that cite critical imaging findings. However, IR tools that have been developed for one disease or imaging modality often need substantial reconfiguration before they can be utilized for another disease entity.
Purpose:
This paper: 1) describes the process of customizing two Natural Language Processing (NLP) and Information Retrieval/Extraction applications – an open-source toolkit, A Nearly New Information Extraction system (ANNIE); and an application developed in-house, Information for Searching Content with an Ontology-Utilizing Toolkit (iSCOUT) – to illustrate the varying levels of customization required for different disease entities and; 2) evaluates each application’s performance in identifying and retrieving radiology reports citing critical imaging findings for three distinct diseases, pulmonary nodule, pneumothorax, and pulmonary embolus.
Results:
Both applications can be utilized for retrieval. iSCOUT and ANNIE had precision values between 0.90-0.98 and recall values between 0.79 and 0.94. ANNIE had consistently higher precision but required more customization.
Conclusion:
Understanding the customizations involved in utilizing NLP applications for various diseases will enable users to select the most suitable tool for specific tasks.
INTRODUCTION
The role of radiological imaging in the diagnosis and treatment of diseases has greatly expanded in the last decade [1-3]. Images are interpreted by radiologists, and pertinent findings recorded in the form of narrative text. In practice, the radiologist should contact the referring clinician whenever a critical imaging result is present. Failure to promptly communicate critical imaging test results is not uncommon and such delays are a major source of malpractice claims in radiology [4-8]. The Joint Commission emphasized a need for improved communication of critical results from diagnostic procedures between and among caregivers by making it a National Patient Safety Goal for 2011 [9].
Our institution established an enterprise-wide communi-cation of Critical Test Results (CCTR) policy for communi-cation of critical imaging results among over 600,000 imaging procedures performed annually [10, 11]. Full implementation of this policy is expected to promote significant safety improvements for a considerable patient population. However, manual analysis to evaluate the policy’s impact on the rate of communicating critical imaging results presented a resource-intensive, time-consuming challenge. A pilot test was conducted on a limited random sample of 12,193 reports from a three year period accounting for approximately 0.7% of the total reports during that time [11]. Although routine evaluation of adherence to the CCTR policy could provide invaluable feedback to end-users (which might be expected to enhance and sustain adherence), continued analysis of a greater number of samples was not sustainable due to the time burden it placed on Radiology division chiefs performing these audits manually.
To overcome this limitation, we used natural language processing and information retrieval/extraction (NLP-IR/E) applications that have been developed for the clinical domain [12-15]. Several NLP applications focus specifically on recognizing clinical findings in radiology reports [16-19]. Evaluation of applications that identify patient information in clinical reports has been done for such foci as smoking status, patient symptoms, and clinical diagnoses, such as psoriatic arthritis and status of tumors [14, 20]. However, these previous evaluations have been limited to single diseases and/or imaging examinations and have not focused on assessing the steps required for customizing or reconfiguring the application for various clinical conditions. Thus, although a system might successfully find cases with documented lung cancer, it may not necessarily be able to find cases with pneumothorax. In many instances, manually reconfiguring an application for a different clinical condition than that for which the tool was developed requires significant effort as well as injection of expert opinion [21]. Describing these customization processes are essential steps for clinical application and performance evaluation [22].
The first objective of this project is to describe the process of customizing two different NLP-IR/E applications – an open-source Information Extraction toolkit utilized by the General Architecture for Text Engineering (GATE) research community called A Nearly New Information Extraction system (ANNIE);[23] and a toolkit developed in-house, named Information for Searching Content with an Ontology-Utilizing Toolkit (iSCOUT) - to retrieve radiology reports with three distinct critical imaging findings (pulmonary nodule, pneumothorax, and pulmonary embolus), illustrating different levels of customization required to use these applications. The second objective is to evaluate application performance in retrieval of radiology reports with the specified critical imaging findings.
MATERIALS AND METHODOLOGY
Study Site and Setting
The study protocol was approved by our institution’s Institutional Review Board. The requirement for obtaining informed consent was waived for this study. Our institution is a 752-bed adult urban tertiary academic medical center with over 600,000 imaging examinations ordered annually.
The Natural Language Processing and Information Retrieval/Extraction (NLP-IR/E) Applications
A description of the applications selected for analysis follows, along with the levels of customization required for this project.
ANNIE - A Nearly New Information Extraction system:
ANNIE is the information extraction component included in GATE, a comprehensive suite of tools utilized by the scientific community for NLP and information extraction and retrieval.(23) Originally developed in 1995 at the University of Sheffield (South Yorkshire, UK), as part of the Large Scale Information Extraction project of the Engineering and Physical Sciences Research Council of the UK, GATE currently has users, developers and contributors from all over the world. ANNIE contains several tools to process textual reports into its basic syntactic components and to run a desired search. The components utilized for this application include: Tokenizer, Sentence Splitter, Part-of-Speech (POS) Tagger, Sectionizer, Gazetteer, and Named Entity (NE) Transducer.
The Tokenizer manipulates the syntactic structure of the narrative, splitting up each sentence by commas, white space, and other forms of punctuation into meaningful elements (e.g., words). This facilitates search for specific tokens. The Sentence Splitter recognizes sentences by reviewing a listing of sentence-ending punctuations and then splits each sentence of a report. The POS Tagger tags each word in a report with its part of speech. This tool relies on a set of rules and a lexicon integrated into the GATE application. The Sectionizer splits reports into sections based on paragraph breaks. Each section is further divided into section headings. For instance, a radiology report would contain the following section headings – Indication, Findings and Impression. The Gazetteer utilizes lists of specific words and phrases as the search criteria and scans through reports, highlighting words and phrases that appear on the gazetteer lists. The textual contents of these lists can be encoded to signify that the report holds positive or negative criteria, based on the programming files supplied by the user. The NE Transducer uses Java Annotation Pattern Engine (jape) files written in jape language to classify the terms that the Gazetteer is programmed to find. For example, the transducer will either negate unrecognized terms or classify as “positive” those matching pre-selected terms in the list files that the Gazetteer utilizes. These tools work collectively to retrieve a set of reports (e.g. reports with unique accession numbers) meeting the user-defined search criteria.
iSCOUT - Information for Searching Content with an Ontology-Utilizing Toolkit:
iSCOUT employs tools that work together to enable a query from a set of unstructured narrative text reports. Various components act to tokenize textual reports into individual words, enable sentence identification, and utilize term matching to implement a query. iSCOUT is comprised of a core set of tools, including the Data Loader, Header Extractor, Terminology Interface, Reviewer and Analyzer. In addition, ancillary components include a Stop Word Remover and a Negator. Using an executable batch file, the components of the toolkit are utilized in series. Similar to ANNIE, all iSCOUT components can be utilized to retrieve reports meeting a user-defined set of criteria. iSCOUT was originally developed at our institution and written in Java programming language [24].
The Report Separator’s main engine is a file separator, which parses the text file containing multiple radiology reports, then formats individual reports, each identified by the report accession number. The Header Extractor limits search to specific parts of the reports, such as findings, impression, and conclusion. The Terminology Interface allows query expansion based on synonymous or other related terms. The Reviewer generates a text file containing the entire report for each of the returned results for a more detailed review. The Analyzer calculates performance metrics, specifically the precision and recall, for a particular query, given a list of accession numbers that should have been retrieved (e.g., gold standard). The Stop Word Remover removes all stop words from the reports while the Negator excludes reports that contain negation terms, such as “no” or “not.” iSCOUT components can be used in combination, selecting individual components as necessary to enable a query and retrieve selected radiology reports.
Radiology Reports
For each of the diseases, radiology reports were randomly selected from tests performed in the Emergency Department from January to June 2010. The target sample sizes were 200 each for chest CT scans and pulmonary embolism (PE) chest CT scans (with a specific protocol designed to look for pulmonary embolism) and 500 reports for chest x-rays, consistent with the relative proportion of ordered tests. For the PE search, a total of only 179 PE chest CT scan reports were available since there were a few duplicates and some regular chest CT scans were misclassified as PE CT reports. For pulmonary nodule, a total of 212 chest CT scan reports were available for analysis. Finally, 500 chest x-ray reports were gathered to search for pneumothorax. Personal health information identifiers were expunged from the reports prior to analysis.
Gold Standard: Reference based on manual review:
A “gold standard” list of radiology reports that reported clinical findings consistent with the diseases being evaluated was generated in order to evaluate the performance of each NLP-IR/E application. The PE chest CT scan reports were manually reviewed by two researchers and one radiologist. Two or more reviewers agreed on positive findings of PE from the textual narrative in 23 of the 179 PE chest CT scan reports. For pulmonary nodule, one researcher and one radiologist reviewed the chest CT scan reports. When both agreed that a report indicated a finding of pulmonary nodule, this report was included in the list. When there was disagreement between the two reviewers, each report was adjudicated by a senior radiologist whose judgment was final. A set of 61 reports was deemed positive for pulmonary nodule. For pneumothorax, 500 chest x-ray reports were reviewed by a researcher and a radiologist who decided by consensus on a total of 31 positive cases. The gold standard sets of accession numbers for each search or query were compared with those generated by each NLP-IR/E application.
Disease-Specific Customization
Each of the applications used in this project was downloaded locally, and utilized for processing radiology reports. The data format required by both applications was a text file containing all reports, delimited by distinct characters in between each report. This pre-processing was a necessary step for every application, and was not identified as a customization task. Similarly, annotation in order to create a gold standard and data analysis were not considered customization tasks.
Several steps are necessary to customize applications for specific diseases. In some cases, the tasks require various types of expertise. Table 1 enumerates and describes various tasks identified for disease-specific customization when utilizing different applications.
Disease-Specific Customization Tasks
| Tasks | No Expertise | Expertise | |
|---|---|---|---|
| Informatics/Programming | Clinical | ||
| Data Entry | X | ||
| Manual Review | X | ||
| Software Development | X | X | |
| Terminology Code Identification | X | X | |
| Feature Selection (e.g. Enumerating Query Lists) | X | X | |
Term Lists for Pulmonary Nodule
| Term List | Terms |
|---|---|
| Positive Term List | spn |
| Negative Term List | no, without, not, cannot, absent, absence, exclude, ruled-out, difficult, unlikely, free, negative, resolved, normal, ground glass opacity, BAC, inflammatory, cyst, thyroid, liver, kidney |
| Combination List A | pulmonary, lung, subpleural, lobe, lobular |
| Combination List B | nodule, nodular opacity |
Term Lists for Pulmonary Embolus
| Term List | Terms |
|---|---|
| Positive Term List | pulmonary embolism, pulmonary embolus, pulmonary emboli, pe, saddle embolism, saddle embolus, saddle emboli |
| Negative Term List | no, without, not, absent, absence, limited, suboptimal, non-diagnostic, exclude, cannot, ruled out, difficult, unlikely, free, negative, never, inadequate, insufficient, resolved, diagnostic, CT, normal, limits, quality, study, resolution, artifact |
| Combination List A | pulmonary |
| Combination List B | filling defect |
Term Lists for Pneumothorax
| Term List | Terms |
|---|---|
| Positive Term List | ptx, pneumothorax, pneumothoraces |
| Negative Term List | no, without, not, absent, absence, negative, resolved |
| Combination List A | pleural space |
| Combination List B | air |
Term Lists for Use with iSCOUT
| Query Subject | Terms |
|---|---|
| Pulmonary Nodule | spn, solitary pulmonary nodule, sub pleural nodule, lung nodule, nodular opacity, pulmonary nodule, pulmonary nodular opacity |
| Pulmonary Embolus | pulmonary embolism, pulmonary embolus, pulmonary emboli, pe, saddle embolism, saddle embolus, saddle embolism, saddle emboli, filling defect |
| Pneumothorax | ptx, pneumothorax, pneumothoraces, pleural space air |
NLP-IR/E Applications and Level of Customization
| Tasks | ANNIE | iSCOUT |
|---|---|---|
| Data Entry | low | low |
| Manual Review | low | low |
| Software Development | high | none |
| Terminology Code Identification | optional | optional |
| Feature Selection (e.g. Enumerating Query Lists) | high | low |
Data entry is necessary to populate each user interface with the appropriate file(s) for processing, as well as to fill query lists or forms with the appropriate search terms. No advanced expertise is necessary to perform this task, and although multiple diseases and imaging modalities typically require increased requirement for data entry, this does not significantly impact task completion. Manual review, on the other hand, requires some technical expertise. This task involves reviewing data files to ensure that they meet the format required by an application. Subsequent re-formatting of the data file or software customization is necessary based on this review. Manual review likewise includes reviewing preliminary results of analysis to ensure that results are not derived erroneously. Unexpected errors may arise due to use of unexpected character delimiters (e.g., blank spaces). In addition, query terms may not be the ones intended. Thus, informatics expertise is often necessary for performing manual review prior to full implementation in a new disease entity or imaging modality. In some instances, clinical expertise may be sought when performing manual review, especially when search terms are ambiguous (i.e. anaphora resolution). However, clinical experts usually do not need to perform manual review. Software development, when necessary, is a potential hurdle when implementing an application. This is particularly true when customization relies heavily on a programmer or a technical expert to perform the task, in conjunction with a clinical expert. Thereafter, the application is augmented with software to enable seamless interface with the analytical file.
The last two tasks included in Table 1 require both technical and clinical expertise. Feature selection, in some cases, can be described as enumerating query lists and is critical in any classification task. Likewise, terminology code identification is often necessary to perform information retrieval and/or extraction. The first task relies on an expert creating lists of terms to find, or lists of terms to avoid, for information retrieval. The second task relies on a controlled terminology to identify codes for appropriate search terms, relying on the terminology to provide other semantically related terms. Either method requires human expertise to appropriately identify necessary terms or codes. Alternatively, feature extraction may be automated, with manual annotation relying on human expertise for identifying documents of interest (e.g. reports containing a finding). In either case, the feature selection process requires both technical and clinical expertise. Similarly, human expertise is required to resolve ambiguities and inconsistencies in the data, as evidenced by those resulting from temporal variability (e.g. “past history” of a finding) or anaphora resolution.
Report Retrieval
ANNIE.
Reformatting of the radiology report text files was completed. In order to utilize the Gazetteer, jape programming files were developed for this project by the investigators. Four different term lists were enabled: a positive term list, a negative term list, and two lists to be used in conjunction – Combination List A and Combination List B.
Table 2 enumerates all the terms listed in the four term lists for the pulmonary nodule query. All lexical variants of the terms (e.g., stem variant, orthographic, order) and plural forms were included in the lists.
The positive term list contains terms, which if found in the text, signify positive findings. As an example, if a sentence contained the word “spn,” the report will be retrieved. The negative term list contains terms which, if found in the text, would flag the report as not having the desired finding. Combination Lists A and B contained words and phrases that also signified positive findings in the reports. However, the two lists’ terms must be seen together in the same sentence for that report to be identified as containing a positive finding. One of these lists, for example, signified areas of the chest (e.g., lung), while the other signified similar terms for “nodule.” Thus, if a sentence contains both the terms “nodule” and “lung,” the report will be marked as having a positive finding.
Tables 3 and 4 enumerate term lists for pulmonary embolus and pneumothorax. Similar algorithm and program files were utilized as those described for pulmonary nodule.
iSCOUT:
As iSCOUT was developed for use at our institution, it did not require further customization or reformatting of the text files. A single term list is required for each query. A Negator is included as one of the components of the toolkit, thus precluding the requirement for a negative term list. Table 5 enumerates the list of terms for each of the queries for pulmonary nodule, pulmonary embolus and pneumothorax. All lexical variants of the terms (e.g., stem variant, orthographic, order) and plural forms were included in the lists. A batch file combines specific components of iSCOUT and returns a list of accession numbers marked as having a positive finding.
Data Analysis
Precision and recall, including confidence intervals, were calculated and recorded for each application [25]. Precision refers to the proportion of true positive (TP) reports from the total number of reports retrieved. Recall refers to the proportion of true positives that were actually retrieved from all reports that should have been retrieved (gold standard). Thus, precision reflects the proportion of reports that were retrieved with the appropriate finding.
RESULTS
The level of customization required for each of the NLP-IR/E applications when applied to multiple diseases is presented in Table 6. Customization is categorized as “high,” when task completion requires performing more than one subtask, leading to greater time and/or effort, and “low,” when there is one subtask required to complete a task. Data entry for ANNIE and iSCOUT requires minimal effort since entire reports are processed in a single input file for both applications. Manual review was “low” for both applications since data files required a single review, leading to an appropriate file format (e.g. character-delimited text files) for data processing. For software development, ANNIE requires jape programming to enable use of ANNIE within GATE, and to run it over a desired corpus (e.g., radiology reports). Jape programming is also required to enable access to the Gazetteer. No further software development was required for iSCOUT. Enumerating query lists is likewise more labor and time-intensive for ANNIE, requiring three lists to be completed for each disease, compared to only a single list for iSCOUT. Terminology code identification, which can be utilized to further expand the list of query terms to enhance report retrieval, was not performed in this study. This task is optional for both applications, and would require further programming for ANNIE, but not for iSCOUT.
The precision and recall for each NLP-IR/E application are shown in Figs. (1-3). Comparison between the three applications is shown for three separate queries for pulmonary nodule, pulmonary embolus and pneumothorax.
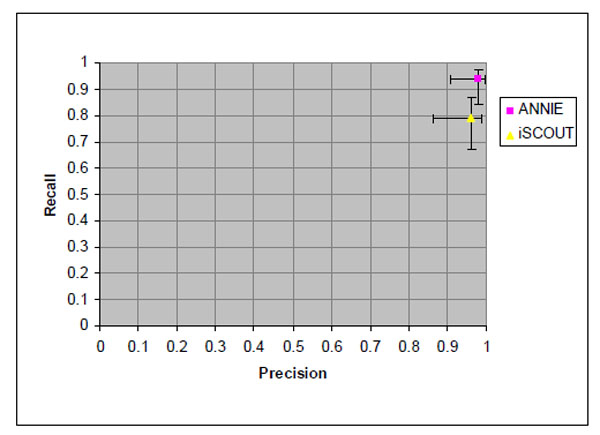 |
Fig. (1). Precision and recall values for retrieving reports with pulmonary nodule and 95% confidence intervals. |
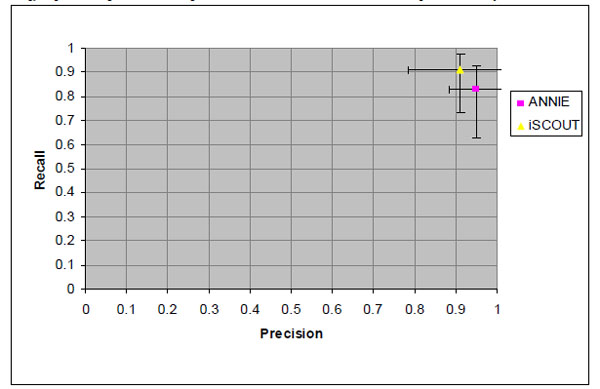 |
Fig. (2). Precision and recall values for retrieving reports with pulmonary embolism and 95% confidence intervals. |
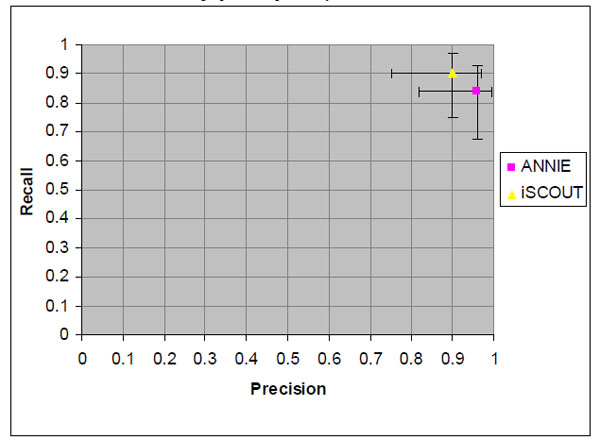 |
Fig. (3). Precision and recall values for retrieving reports with pneumothorax and 95% confidence intervals. |
Pulmonary Nodule
Fig. (1) illustrates the precision and recall for both NLP-IR/E applications. ANNIE had a higher precision and recall at 0.98 and 0.94, respectively. iSCOUT had a precision of 0.96 and a recall of 0.79. However, the differences were not statistically significant. The 95% confidence limits for each value are indicated separately for precision and recall according to the matrix axis.
Pulmonary Embolus
Fig. (2) shows the precision and recall values for the PE query, similarly with 95% confidence intervals marked along each axis. ANNIE demonstrated higher precision at 0.95, with a recall of 0.83. iSCOUT showed higher recall at 0.91, with precision also at 0.91.
Pneumothorax
Demonstrated in Fig. (3) are the precision and recall values for the pneumothorax query, again with 95% confidence intervals marked along each axis. The NLP-IR/E applications collectively performed well for precision at 0.96 (ANNIE) and 0.90 (iSCOUT). Recall was higher for iSCOUT at 0.90 compared to ANNIE at 0.84.
DISCUSSION
This project describes two NLP-IR/E applications and compares their performance in retrieving radiology reports that describe three distinct critical findings. Unlike other previous evaluations, this project focuses on both customization as well as accuracy of these applications for disease-specific retrieval. Customization of an application involves restructuring a system’s functionality to accommodate specific goals. Customization requires close coordination with experts, both in clinical as well as technical matters. Thus, a project’s goals as well as available resources may dictate which application is most appropriate, assuming accuracy measures of the applications are comparable.
This evaluation focuses on three distinct clinical diseases. However, the applications are expected to be utilized for retrieving reports with many more critical imaging findings. Retrieving reports with pulmonary nodule can provide insight into the rate of follow-up for incidental pulmonary nodules and determine if it is consistent with the Fleischner Criteria, as recommended in a national clinical practice guideline [26]. Identifying and retrieving reports that describe findings consistent with pulmonary embolism is important for assessing quality of patient care [27]. Quality assessment of CT orders and ordering patterns for suspected pulmonary embolism is vital to reduce unnecessary radiation exposure [3]. For pneumothorax, the ability to retrieve specific reports can provide insight into an investigation of adverse events related to central line placements, as pneumothorax is a known complication of this procedure [28, 29]. More importantly, the ability to automatically identify reports with any of these three critical findings can provide a reminder for radiologists to alleviate delays in communicating critical results. In addition, it can be used to routinely select cases in order to determine prevalence rates of timely communication of critical results, measuring adherence to a CCTR policy and allowing for continuous quality assurance and process improvement initiatives.
With regard to the precision and recall of each application, ANNIE and iSCOUT both consistently displayed similar excellent precision and recall values [19, 20, 30]. Overlap of 95% confidence intervals for each application indicates no significant differences in precision and recall between the two. These results are not unexpected given the similarity in NLP-IR/E algorithms underlying these applications.
ANNIE attained higher precision values for all three queries. The ability to reduce false positives is due largely to its ability to maintain unique negation lists for specific searches. The negation lists, in particular, helps refine searches to only specific instances where pulmonary nodules are seen. For example, we would want to negate a report that indicates an attenuating nodule that “likely represents a sebaceous cyst”. The ability to include the word “cyst” in the negation list mitigates this problem. We were able to do this with a number of other terms, including “liver” and “kidney”to focus the query only to the lungs. This increased precision, however, come at a cost. ANNIE needs to be manually programmed to the end user’s specifications in order to accommodate and access specific lists. In addition, selecting terms that are included in each of the four potential lists is not an easy task, requiring careful thought from one or more experts. Reviewing local term usage from sample reports a priori becomes a critical component for identifying appropriate terms that will be included in each list.
iSCOUT demonstrated high precision and recall, similar to ANNIE. Utilizing the Header Extractor significantly increased precision, since the search was limited to certain sections of the report. For example, the phrase, “Multiple axial CT images of the chest were obtained following IV contrast administration using a pulmonary embolism protocol,” contains the term “pulmonary embolism” and would have caused the tools to erroneously retrieve reports with this protocol stated in the Technique section of the report. The ease of performing a query utilizing iSCOUT was significant, given that no additional programming is required. In addition, populating a single term list was conveniently simple. For pneumothorax, for instance, the term list only had four terms (see Table 5).
LIMITATIONS
This retrospective study compared the use of two applications for retrieving reports with critical imaging findings from a single institution. Further testing in multiple settings would document generalizability of both applications for various clinical domains. Moreover, retrieval was based on documented presence of the findings within textual reports. Thus, documentation of findings is essential in evaluating information retrieval.
In addition, both applications were utilized to maximize the following metrics for information retrieval - precision and recall. Feature selection (e.g. enumerating query lists) was performed empirically, and not exhaustively. Utilizing controlled terminology could have further increased recall, as previously demonstrated in retrieving liver cysts using iSCOUT [24]. Finally, supervised classification algorithms, previously implemented for information retrieval, were not available in either application [31]. Incorporating these algorithms into information retrieval applications could further enhance precision and recall of these tools.
CONCLUSION
Both applications can be used for information retrieval, comparing favorably with several information retrieval tools currently in use [19, 30-32]. ANNIE had consistently high precision values, although using it required the most customization. iSCOUT performed equally well with less customization. Appropriate utilization of these applications for retrieving reports with critical findings is important in ensuring adherence to practice guidelines. The American College of Radiology (ACR) in its 2005 Practice Guideline for Communication of Diagnostic Imaging Findings strongly advised radiologists to expedite notification of imaging results to referring physicians “in a manner that reasonably ensures timely receipt of the findings” [33]. Utilizing these applications in retrieving reports with critical findings and verifying whether timely and appropriate communication was initiated can greatly expedite the evaluation process and provide invaluable feedback to caregivers.
ACKNOWLEDGEMENTS
This work was funded in part by Grant R18HS019635 from the Agency for Healthcare Research and Quality.
CONFLICT OF INTEREST
The author confirms that this article content has no conflict of interest.
REFERENCES
| [1] | Sunshine JH, Hughes DR, Meghea C, Bhargavan M. What Factors Affect the Productivity and Efficiency of Physician Practices? Med Care 2010; 48(2 ): 110-7. |
| [2] | Swayne LC. The private-practice perspective of the manpower crisis in radiology: greener pastures? J Am Coll Radiol 2004; 1(11 ): 834-41. |
| [3] | Sodickson A, Baeyens PF, Andriole KP, et al. Recurrent CT, cumulative radiation exposure, and associated radiation-induced cancer risks from CT of adults Radiology 2009; 251(1 ): 175-84. |
| [4] | Gordon JR, Wahls T, Carlos RC, Pipinos II, Rosenthal GE, Cram P. Failure to recognize newly identified aortic dilations in a health care system with an advanced electronic medical record Ann Intern Med 2009; 151(1 ): 21-7.: W5. |
| [5] | Singh H, Arora HS, Vij MS, Rao R, Khan MM, Petersen LA. Communication outcomes of critical imaging results in a computerized notification system J Am Med Inform Assoc 2007; 14(4 ): 459-66. |
| [6] | Roy CL, Poon EG, Karson AS, et al. Patient safety concerns arising from test results that return after hospital discharge Ann Intern Med 2005; 143(2 ): 121-8. |
| [7] | Johnson CD, Krecke KN, Miranda R, Roberts CC, Denham C. Quality initiatives: developing a radiology quality and safety program: a primer Radiographics 2009; 29(4 ): 951-. |
| [8] | Berlin L. Statute of limitations and the continuum of care doctrine AJR Am J Roentgenol 2001; 177(5 ): 1011-6. |
| [9] | The Joint Commission. 2011 National Patient Safety Goals. Available from: http://www.jointcommission.org/standards_inform ation/npsgs.aspx 2011 [Cited: 2012 Jan 11]; |
| [10] | Hunsaker A, Khorasani R, Doubilet P. Policy for Communicating Critical and/or Discrepant Results Available from: http://www. brighamandwomens.org/cebi/files/Radiology%20Policy%20Critica l-Discrepant%20 Results.pdf. 3-10-2009 [Cited: 2012 Jan 11]; |
| [11] | Khorasani R. Optimizing communication of critical test results J Am Coll Radiol 2009; 6(10 ): 721-3. |
| [12] | Friedman C, Alderson PO, Austin JH, Cimino JJ, Johnson SB. A general natural-language text processor for clinical radiology J Am Med Inform Assoc 1994; 1(2 ): 161-74. |
| [13] | Zingmond D, Lenert LA. Monitoring free-text data using medical language processing Comput Biomed Res 1993; 26(5 ): 467-81. |
| [14] | Zeng QT, Goryachev S, Weiss S, Sordo M, Murphy SN, Lazarus R. Extracting principal diagnosis, co-morbidity and smoking status for asthma research: evaluation of a natural language processing system BMC Med Inform Decis Mak 2006; 6: 30. |
| [15] | Savova GK, Masanz JJ, Ogren PV, et al. Mayo clinical Text Analysis and Knowledge Extraction System (cTAKES): architecture, component evaluation and applications J Am Med Inform Assoc 2010; 17(5 ): 507-13. |
| [16] | Taira RK, Soderland SG, Jakobovits RM. Automatic structuring of radiology free-text reports Radiographics 2001; 21(1 ): 237-45. |
| [17] | Mamlin BW, Heinze DT, McDonald CJ. Automated extraction and normalization of findings from cancer-related free-text radiology reports AMIA Annu Symp Proc 2003; 420-. |
| [18] | Thomas BJ, Ouellette H, Halpern EF, Rosenthal DI. Automated computer-assisted categorization of radiology reports AJR Am J Roentgenol 2005; 184(2 ): 687-90. |
| [19] | Dreyer KJ, Kalra MK, Maher MM, et al. Application of recently developed computer algorithm for automatic classification of unstructured radiology reports: validation study Radiology 2005; 234(2 ): 323-9. |
| [20] | Meystre S, Haug PJ. Natural language processing to extract medical problems from electronic clinical documents: performance evaluation J Biomed Inform 2006; 39(6 ): 589-99. |
| [21] | Dixon BE, Samarth A. Innovations in Using Health IT for Chronic Disease Management AHRQ National Resource Center for Health Information Technology AHRQ Publication No 09-0029-EF 2009. |
| [22] | Reiner BI. Customization of medical report data J Digit Imaging 2010; 23(4 ): 363-73. |
| [23] | Cunningham H, Maynard D, Bontcheva K, Tablan V. GATE: A framework and graphical development environment for robust NLP tools and applications In: Proceeding of the 40th Anniversary Meeting of the Association for Computations Lingusitics (ACL’02); July 2002; Philadelphia. |
| [24] | Lacson R, Andriole KP, Prevedello LM, Khorasani R. Information from Searching Content with an Ontology-Utilizing Toolkit (iSCOUT) J Digit Imaging 2012; 25(4 ): 512-9. |
| [25] | Hersh W. Evaluation of biomedical text-mining systems: lessons learned from information retrieval Brief Bioinform 2005; 6(4 ): 344-56. |
| [26] | MacMahon H, Austin JH, Gamsu G, et al. Guidelines for management of small pulmonary nodules detected on CT scans: a statement from the Fleischner Society Radiology 2005; 237(2 ): 395-400. |
| [27] | Duriseti RS, Brandeau ML. Cost-effectiveness of strategies for diagnosing pulmonary embolism among emergency department patients presenting with undifferentiated symptoms Ann Emerg Med 2010; 56(4 ): 321-. |
| [28] | Harley KT, Wang MD, Amin A. Common procedures in internal medicine: improving knowledge and minimizing complications Hosp Pract (Minneap) 2009; 37(1 ): 121-7. |
| [29] | Ayas NT, Norena M, Wong H, Chittock D, Dodek PM. Pneumothorax after insertion of central venous catheters in the intensive care unit: association with month of year and week of month Qual Saf Health Care 2007; 16(4 ): 252-5. |
| [30] | Cheng LT, Zheng J, Savova GK, Erickson BJ. Discerning tumor status from unstructured MRI reports--completeness of information in existing reports and utility of automated natural language processing J Digit Imaging 2010; 23(2 ): 119-32. |
| [31] | D'Avolio LW, Nguyen TM, Farwell WR, et al. Evaluation of a generalizable approach to clinical information retrieval using the automated retrieval console (ARC) J Am Med Inform Assoc 2010; 17(4 ): 375-82. |
| [32] | Savova GK, Fan J, Ye Z, et al. Discovering peripheral arterial disease cases from radiology notes using natural language processing AMIA Annu Symp Proc 2010 2010; 722-6. |
| [33] | Kushner DC, Lucey LL. Diagnostic radiology reporting and communication: the ACR guideline J Am Coll Radiol 2005; 2(1 ): 15-21. |



